Introduction
In this vignette, we will walk through transittraj’s
entire AVL cleaning workflow, intended to correct or remove problematic
point observations and trips. This seven-step process is summarized in
the figure below.
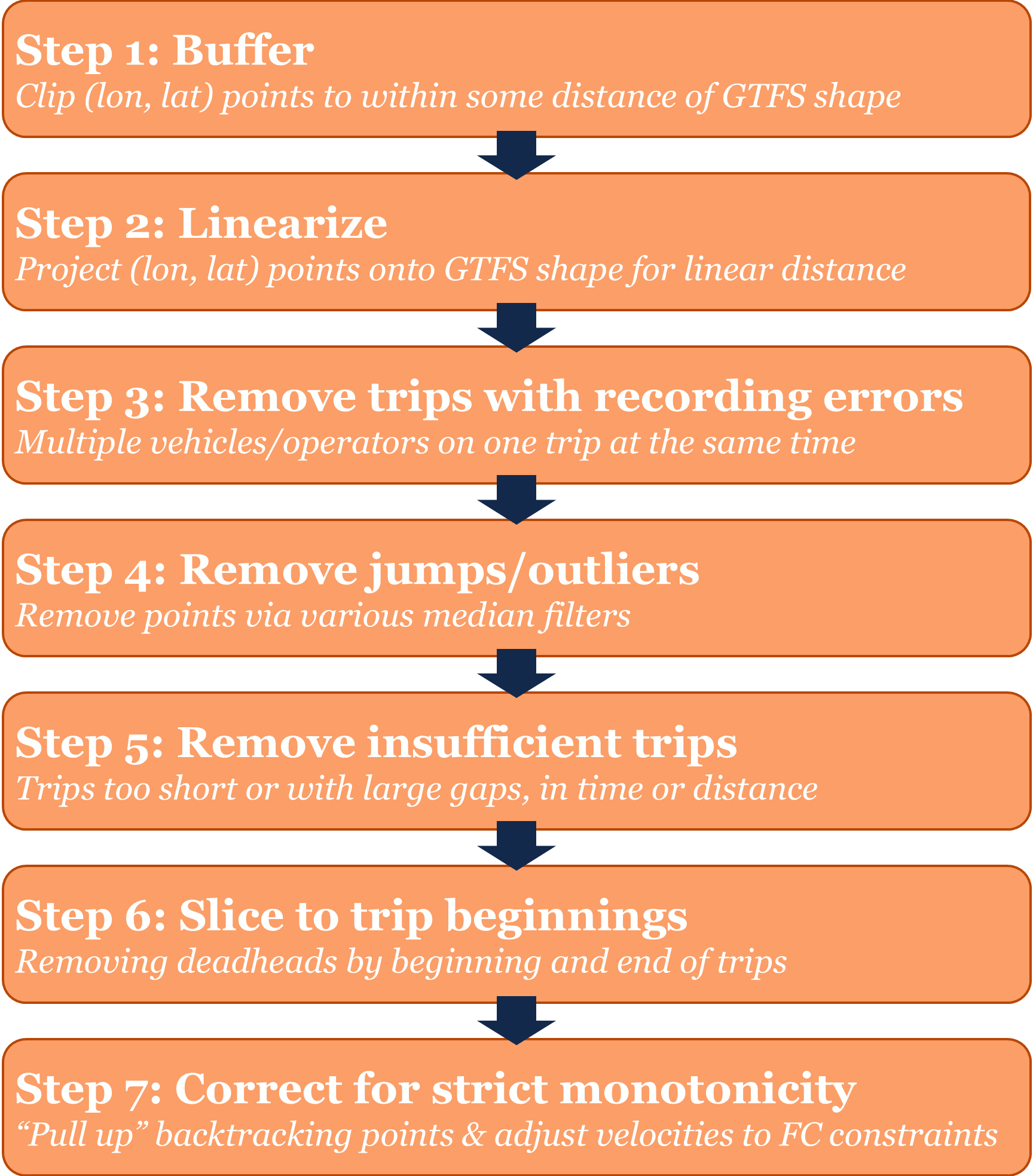
transittraj AVL cleaning process
Check out vignette("input-data") to learn more about the
AVL and GTFS data we’ll be using. Let’s begin by loading the libraries
we’ll be using:
Step 0: Setup
We’ll begin by setting up our data. We’ll be using
wmata_avl and wmata_gtfs for our TIDES AVL and
static GTFS input, respectively.
AVL
For this example, we’ll work with WMATA’s C53 bus route in the northbound direction. Let’s filter our AVL data to these specifications:
# Set our filtering parameters
c53 <- "C53"
c53_dir <- 0 # 0 is NB, 1 is SB
# Filter the entire AVL to just the NB C53
c53_avl <- wmata_avl %>%
filter((route_id == c53) & (direction_id == c53_dir))Now our dataset should include only northbound C53 trips. We can see below that this window has about 5,400 individual GPS pings across 33 trips and 5 hours.
# Pull attributes about our filtered AVL
total_obs <- dim(c53_avl)[1]
num_trips <- length(unique(c53_avl$trip_id_performed))
time_span <- round(max(c53_avl$event_timestamp) - min(c53_avl$event_timestamp),
2)
# Print attributes
cat("Total Observations: ", total_obs,
"\nNumber of trips: ", num_trips,
"\nTime span: ", time_span, " hr",
sep = "")
#> Total Observations: 5436
#> Number of trips: 33
#> Time span: 5.02 hrGTFS
Next, we should grab the GTFS shape and feed information we want for
this route and direction. transittraj provides a couple
functions to help with this. We’ll start by filtering our GTFS to just
the northbound C53:
# Filter the entire GTFS down to NB C53
c53_gtfs <- filter_by_route(gtfs = wmata_gtfs,
route_ids = c53,
dir_id = c53_dir)If we print a summary of this GTFS object, we’ll see that it only contains one route:
summary(c53_gtfs)
#> tidygtfs object
#> files agency, routes, stop_times, trips, shapes, calendar, calendar_dates, stops
#> agency WMATA
#> service from 2025-12-14 to 2026-06-13
#> uses stop_times (no frequencies)
#> # routes 1
#> # trips 952
#> # stop_ids 57
#> # stop_names 57
#> # shapes 2Now that we have the filtered GTFS, we must pull the shape we want as a simple feature (SF) spatial object. You will need to know two things to do this:
Your
shape_id. As shown above, there are twoshape_ids assigned to the northbound C53. Check outplot_interactive_gtfs()to explore each shape individually and decide which one is correct for you. Here we useshape_id = "C53:04".A coordinate reference system. We recommend projecting all spatial data into an appropriate Euclidian coordinate system. Below we use the UTM zone for Washington, DC, Zone 18N (with an EPSG number of 32618).
Now we can use get_shape_geometry() to grab the
shape_id we want, turn it into a spatial line, and project
it to the appropriate coordinate system:
# Set our parameters
c53_NB_shape_id <- "C53:04"
dc_CRS <- 32618
# Pull the shape we want
c53_shape <- get_shape_geometry(gtfs = c53_gtfs,
shape = c53_NB_shape_id,
project_crs = dc_CRS)If we print this new object, we’ll see that it is one multilinestring
with shape_id = "C53:04" and the coordinate system UTM
18N.
print(c53_shape)
#> Simple feature collection with 1 feature and 1 field
#> Geometry type: MULTILINESTRING
#> Dimension: XY
#> Bounding box: xmin: 322061.8 ymin: 4301418 xmax: 329233.3 ymax: 4310353
#> Projected CRS: WGS 84 / UTM zone 18N
#> # A tibble: 1 × 2
#> shape_id geometry
#> <chr> <MULTILINESTRING [m]>
#> 1 C53:04 ((327507.1 4301484, 327505.6 4301478, 327504.9 4301473, 327504 43014…Visualizing our Starting Data
We now have all the data we need to proceed. But before we do, let’s visualize the AVL points and route geometry so we understand what we’re working with:
# Convert GPS points to spatial objects
# You typically won't need to do this -- this is just for visualization
c53_sf <- c53_avl %>%
# As SF
st_as_sf(coords = c("longitude", "latitude"),
crs = 4326) %>%
# Project to DC
st_transform(crs = dc_CRS)
# Generate a map
avl_map <- ggplot() +
# Basemap from OSM
ggspatial::annotation_map_tile(type = "cartolight", zoomin = 0,
progress = "none") +
# Add alignment & points
geom_sf(data = c53_shape, color = "red", linewidth = 1.5) +
geom_sf(data= c53_sf, color = "blue",
alpha = 0.2, size = 0.4) +
# Format our map
theme_void() +
labs(title = "C53 Shape & AVL") +
theme(text = ggplot2::element_text(size = 3))
avl_map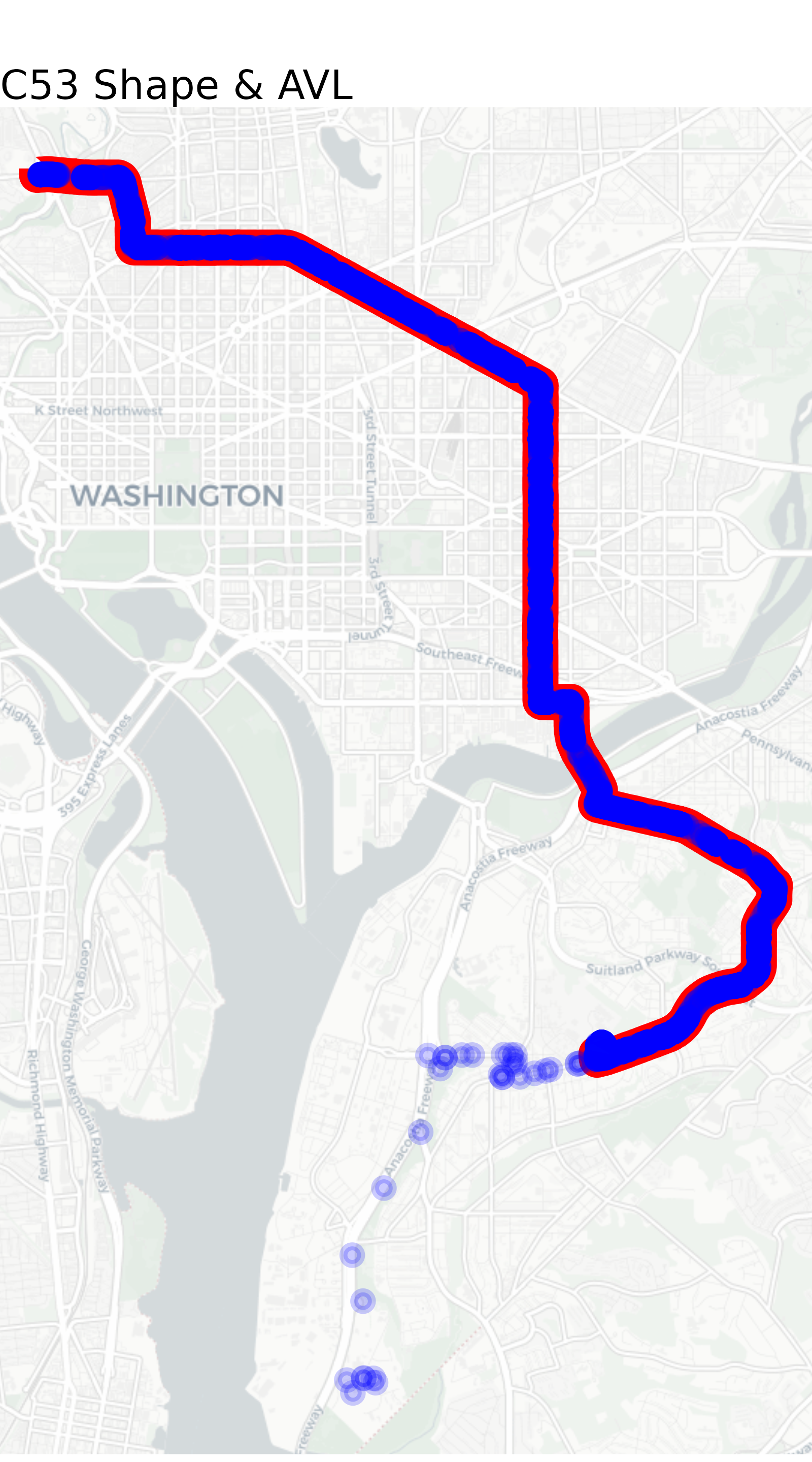
For the most part, our GPS pings follow the route alignment
excellently. But what’s the deal with the points off-route in the far
south? These pings roughly follow I-295 down to WMATA’s Shepherd Parkway
bus garage. These points most likely correspond to one or more
deadheading vehicle that were logged into a trip but were not actually
in revenue service. transittraj includes functions intended
to identify and remove potential deadheads. We’ll begin exploring these
in the following section.
Steps 1 & 2: Buffer & Linearize
Now that we have our data, we can begin cleaning and processesing it. The first two steps we recommend are:
Step 1: Buffer. Clip GPS points to within a certain distance of the route alignment.
Step 2: Linearize. Project GPS points onto the route alignment to retrieve the linear distance from the start of the route.
Both steps are bundled into one function,
get_linear_distances(). There are two main decision
variables for this function:
The projection CRS. This should be the same one you used to create your route shape.
The buffer distance. This will be in the units of your spatial projection. Here we’re using UTM, which is in meters. Thus, all distance measurements you see in the vignette will be in meters.
For this example, we’ll continue using UTM Zone 18N, and we’ll only keep points within 50 meters of the route alignment. This should take care of those deadheading points we saw earlier.
Running the Code
We can now clip and linearize our AVL dataframe using
get_linear_distances():
# Set parameters
c53_buffer = 50 # meters
# Run the cleaning function
c53_distances <- get_linear_distances(avl_df = c53_avl,
shape_geometry = c53_shape,
project_crs = dc_CRS,
clip_buffer = c53_buffer)Exploring the Results
Now each observation is represented by a distance from the route’s beginning. Let’s first see how many observations were clipped out:
# Pull dimensions of each
step0_obs <- dim(c53_avl)[1]
step1_obs <- dim(c53_distances)[1]
# Print
cat("Initial: ", step0_obs, " obs",
"\nAfter buffer: ", step1_obs, " obs",
"\nDifference: ", (step0_obs - step1_obs), " obs removed")
#> Initial: 5436 obs
#> After buffer: 4946 obs
#> Difference: 490 obs removedWe had just under 500 observations clipped out of our dataset. We can also take a look at the new header of our dataset:
head(c53_distances)
#> location_ping_id vehicle_id trip_id_performed service_date route_id
#> 1 1 5461 18632100 2026-02-16 C53
#> 2 3 5464 14078100 2026-02-16 C53
#> 3 4 5466 25836100 2026-02-16 C53
#> 4 6 5473 8428100 2026-02-16 C53
#> 6 8 5481 1115100 2026-02-16 C53
#> 7 9 5479 842100 2026-02-16 C53
#> direction_id speed trip_stop_sequence event_timestamp stop_id
#> 1 0 0.0000 63 2026-02-16 10:58:09 7219
#> 2 0 10.9728 52 2026-02-16 10:58:31 6843
#> 3 0 0.0000 2 2026-02-16 10:58:27 13111
#> 4 0 4.5720 63 2026-02-16 10:58:02 7219
#> 6 0 7.9248 41 2026-02-16 10:58:10 28286
#> 7 0 8.8392 29 2026-02-16 10:58:24 4520
#> distance
#> 1 1.534625e+04
#> 2 1.205652e+04
#> 3 4.191312e-02
#> 4 1.535750e+04
#> 6 9.512327e+03
#> 7 6.551790e+03Now, instead of latitude and longitude
columns, we have a single numeric distance column. We will
only work with these one-dimensional distances for the rest of this
vignette.
Step 3: Cleaning Overlapping Subtrips
In some AVL systems, multiple vehicle or operator IDs can be recorded
for the same trip. Sometimes this is okay: there may, for example, be an
operator shift change mid-trip. Other times, however, these multiple
vehicles/operators are logged in at the same time (“overlapping”). This
creates problems, as it becomes impossible for transittraj
to understand what the trip is supposed to be doing.
For Step 3 of the cleaning workflow,
clean_overlapping_subtrips() identifies and removes these
trips. Each distinct “subtrip” (a unique combination of trip ID, vehicle
ID, and operator ID) is identified, then we check whether the time
ranges of these subtrips overlap. There are a few key decision
variables:
Should operator IDs be checked? This will look for overlapping operator IDs in each trip.
If a subtrip has only one observation assigned to it, should it be removed?
Should trips with multiple subtrips that do not overlap be removed?
The WMATA dataset does not have include operator IDs, so we will not check for operators. We’ll say that it’s okay to leave in subtrips that aren’t overlapping.
Running the Code
Below we use c53_cleaned_subtrips() to identify and
remove problematic trips:
# Set parameters
c53_check_op <- FALSE
c53_remove_singles <- TRUE
c53_remove_non_overlap <- FALSE
# Run function
c53_cleaned_subtrips <- clean_overlapping_subtrips(
distance_df = c53_distances,
check_operator = c53_check_op,
remove_single_observations = c53_remove_singles,
remove_non_overlapping = c53_remove_non_overlap
)Exploring the Results
Let’s see how many trips or observations were removed:
# Pull new dimensions
step3_obs <- dim(c53_cleaned_subtrips)[1]
# Print
cat("Initial: ", step1_obs, " obs",
"\nAfter: ", step3_obs, " obs",
"\nDifference: ", (step1_obs - step3_obs), " obs removed")
#> Initial: 4946 obs
#> After: 4945 obs
#> Difference: 1 obs removedWe can see that only one observation violated our requirements. To
see what that observation is, we’ll introduce a new feature of
transittraj: the return_removals parameter.
Each cleaning function used in this vignette allows a parameter
return_removals that, when set to TRUE, will
return a dataframe of the points/trips removed through that step and a
brief explanation of why they were removed. Let’s check it out:
# Get removals from previous function
c53_step3_removals <- clean_overlapping_subtrips(
# Same settigns as before
distance_df = c53_distances,
check_operator = c53_check_op,
remove_single_observations = c53_remove_singles,
remove_non_overlapping = c53_remove_non_overlap,
# Return removals
return_removals = TRUE
)
# Print removed point
print(c53_step3_removals)
#> # A tibble: 1 × 7
#> trip_id_performed subtrip n_obs reason action time_range n_subtrips_in_range
#> <chr> <chr> <int> <chr> <chr> <iv<dbl>> <int>
#> 1 8428100 8428100-… 1 singl… remov… [NA, NA) NAWe can see the trip and subtrip
(trip_id-vehicle_id) which was removed, and
that it was removed because it was a single observation. Let’s see what
this trip looked like in the original dataset:
# Filter & print to violating trip
print(c53_distances %>%
filter(trip_id_performed == "8428100"))
#> location_ping_id vehicle_id trip_id_performed service_date route_id
#> 1 6 5473 8428100 2026-02-16 C53
#> direction_id speed trip_stop_sequence event_timestamp stop_id distance
#> 1 0 4.572 63 2026-02-16 10:58:02 7219 15357.5This trip, in fact, only had one observation total. It makes sense to remove this – we can’t do much with a single observation.
Step 4: Clean Outlier/Jumps
GPS data, especially in urban areas, is noisy. Sometimes that noise
manifests as a large instantaneous jump that does not match an points’s
surrounding observations. In Step 4, the function
clean_jumps() detects and removes these using median
filters. This has two main decision variables:
The neighborhood width, the total number of points to consider around each observation.
The maximum distance between a point and the median of the neighborhood around it. This can be defined as an absolute distance from the median, in distance units, or as a multiple of the window’s median absolute deviation (MAD) (this is known as a Hampel filter).
For this example, we’ll use the default neighborhood width of 7 (3 points on either side of each observation). We’ll keep our outlier criteria simple (though not very robust) and use a maximum deviation of 20 meters. This means that if any given point is more than 20 meters away from the median of its surrounding points, it will be tossed.
Running the Code
Let’s run clean_jumps() as discussed above. The only
input we need to change from its default is the maximum distance
deviation.
# Set parameters
c53_max_jump <- 20 # meters
c53_min_jump <- -1 * c53_max_jump # meters
# Run function
c53_no_jumps <- clean_jumps(distance_df = c53_cleaned_subtrips,
max_median_deviation = c53_max_jump,
min_median_deviation = c53_min_jump,
t_cutoff = Inf)Exploring the Results
Let’s see how many points were removed as outliers from this filter:
# Pull dimensions
step4_obs <- dim(c53_no_jumps)[1]
# Print
cat("Initial: ", step3_obs, " obs",
"\nAfter: ", step4_obs, " obs",
"\nDifference: ", (step3_obs - step4_obs), " obs removed")
#> Initial: 4945 obs
#> After: 4934 obs
#> Difference: 11 obs removedWith these settings, 11 points were removed. We can use
return_removals = TRUE, just like in Step 3, to take a
closer look at these outlying points:
# Get removals
c53_step4_removals <- clean_jumps(
# Same settings as before
distance_df = c53_cleaned_subtrips,
max_median_deviation = c53_max_jump,
min_median_deviation = c53_min_jump,
t_cutoff = Inf,
# Return removals
return_removals = TRUE)
# Print the removed points
print(c53_step4_removals)
#> # A tibble: 11 × 13
#> trip_id_performed event_timestamp distance location_ping_id window_med
#> <chr> <dttm> <dbl> <chr> <dbl>
#> 1 1306100 2026-02-16 12:43:47 4954. 16770 4884.
#> 2 1306100 2026-02-16 12:43:57 4954. 16798 4896.
#> 3 1306100 2026-02-16 12:44:05 4844. 16826 4936.
#> 4 1306100 2026-02-16 12:44:35 4896. 16910 4954.
#> 5 1306100 2026-02-16 13:31:22 13819. 24579 13789.
#> 6 13478100 2026-02-16 15:37:40 108. 45428 66.7
#> 7 35294100 2026-02-16 11:28:14 83.2 4721 20.1
#> 8 35294100 2026-02-16 11:32:14 0.0419 5369 20.1
#> 9 35294100 2026-02-16 11:37:49 0.0419 6260 20.1
#> 10 35294100 2026-02-16 11:39:13 257. 6476 83.2
#> 11 35294100 2026-02-16 11:40:34 11.2 6692 214.
#> # ℹ 8 more variables: window_mad <dbl>, med_dist <dbl>, is_implosion <lgl>,
#> # is_tail <lgl>, ignore_observation <lgl>, mad_ok <lgl>, dev_ok <lgl>,
#> # all_ok <lgl>Let’s explore pings 16770 through 16910 from trip 1306100. We can plot the trip in this area to visualize the outliers:
# Filter dataframe to our tirp & distances
plot_df <- c53_cleaned_subtrips %>%
filter(trip_id_performed == "1306100") %>%
filter((distance >= 4500) & (distance <= 5500)) %>%
# Join removals
left_join(y = (c53_step4_removals %>% select(location_ping_id, all_ok)),
by = "location_ping_id") %>%
mutate(all_ok = tidyr::replace_na(all_ok, TRUE))
# Create a plot
jumps_plot <- ggplot() +
# Plot the points
geom_point(data = plot_df,
aes(x = event_timestamp, y = distance,
color = all_ok, shape = all_ok),
size = 3, stroke = 3) +
# Format the points
scale_color_manual(name = "Point OK?",
values = c("FALSE" = "firebrick",
"TRUE" = "darkgreen")) +
scale_shape_manual(name = "Point OK?",
values = c("FALSE" = 4,
"TRUE" = 16)) +
# Format the plot
theme_minimal() +
labs(x = "Time",
y = "Distance (m)",
title = "Outliers on C53",
subtitle = "Trip 13478100")
jumps_plot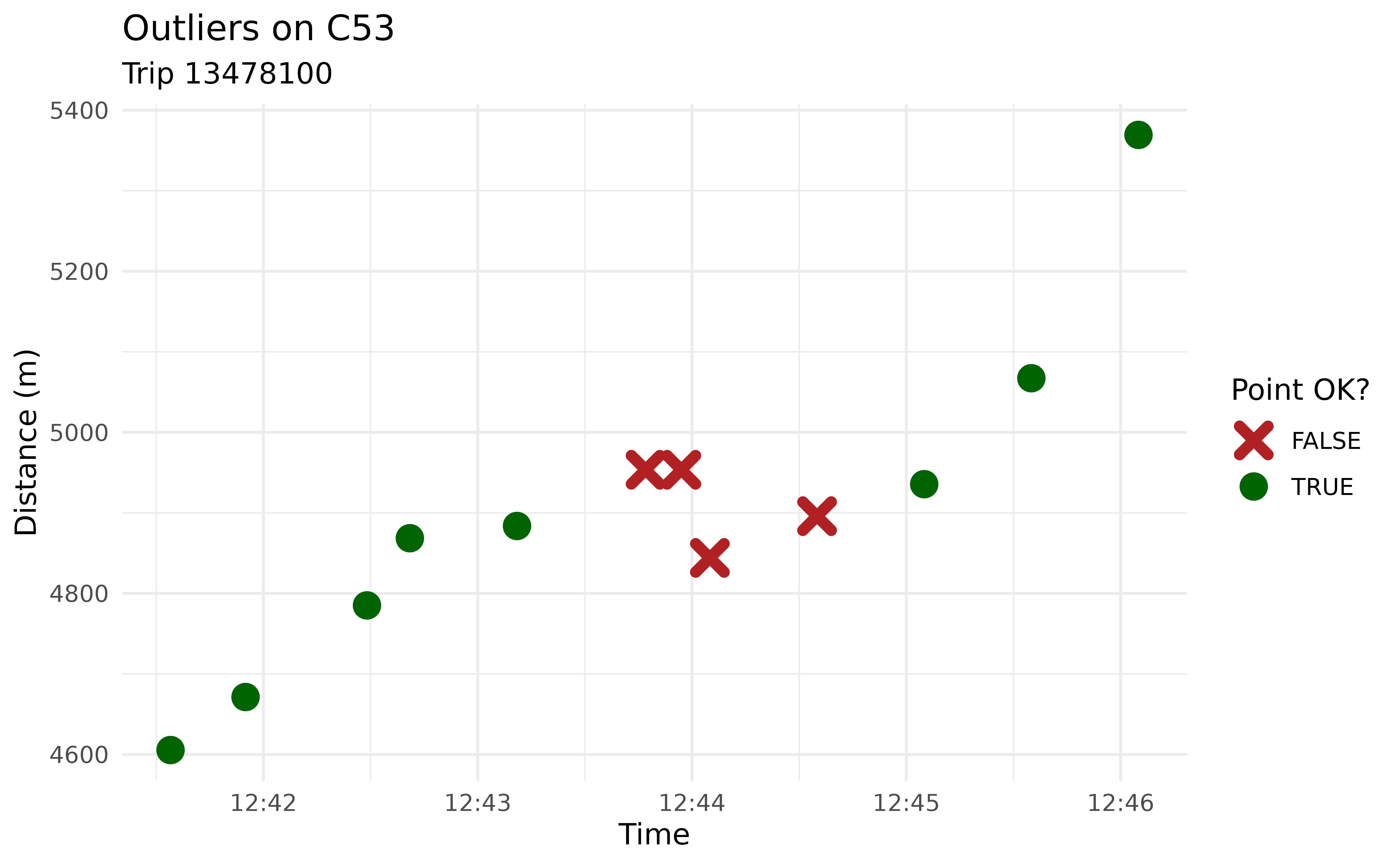
This plot makes it pretty clear that some of these points are a bit
out of line. The final one removed may not truly be an outlier; we
recommend playing around with the function’s options to find settings
that make sense for your data. Read more about these options and the
theory behind median filters at help(clean_jumps).
Step 5: Clean Incomplete Trips
AVL or GTFS-rt data is rarely transmitted perfectly. Often, there may
be large gaps of missing data in the middle of trips, or you may have
only a few observations from each trip. For Step 5, the
function clean_incomplete_trips() filters out these trips
There are two main groups of decision variables for this function:
Minimum and maximum trip distances and durations.
Minimum and maximum distance and time gaps between adjacent observations.
For this example, we’ll use a minimum trip distance of 500 meters, and a minimum trip duration of 90 seconds. Anything will less data than this will be filtered out. Additionally, we’ll remove trips with a gap larger than 500 meters.
Running the Code
Let’s run clean_incomplete_trips() as discussed
above:
# Set parameters
c53_min_dist <- 500 # meters
c53_min_time <- 90 # seconds
c53_max_gap <- 500 # meters
# Run function
c53_cleaned_incompletes <- clean_incomplete_trips(
distance_df = c53_no_jumps,
min_trip_distance = c53_min_dist,
min_trip_duration = c53_min_time,
max_distance_gap = c53_max_gap
)Exploring the Results
Now that we’ve filtered our dataset, let’s see how many trips have been removed:
step5_obs <- dim(c53_cleaned_incompletes)[1]
step5_trips <- length(unique(c53_cleaned_incompletes$trip_id_performed))
step4_trips <- length(unique(c53_no_jumps$trip_id_performed))
cat("Initial: ", step4_obs, " obs, ", step4_trips, " trips",
"\nAfter: ", step5_obs, " obs, ", step5_trips, " trips",
"\nDifference: ", (step4_obs - step5_obs), " obs, ",
(step4_trips - step5_trips), " trips removed",
sep = "")
#> Initial: 4934 obs, 31 trips
#> After: 4089 obs, 24 trips
#> Difference: 845 obs, 7 trips removed7 trips violated our requirements, corresponding to roughly 850
individual observations. As before, we can use
return_removals to take a look at the violating trips:
c53_step5_removals <- clean_incomplete_trips(
# Same settings as before
distance_df = c53_no_jumps,
min_trip_distance = c53_min_dist, min_trip_duration = c53_min_time,
max_distance_gap = c53_max_gap,
# Return removals
return_removals = TRUE
)
print(c53_step5_removals)
#> # A tibble: 7 × 16
#> trip_id_performed max_dist min_dist max_time min_time
#> <chr> <dbl> <dbl> <dttm> <dttm>
#> 1 13927100 127. 0 2026-02-16 14:36:57 2026-02-16 14:36:27
#> 2 14639100 15362. 0 2026-02-16 12:38:36 2026-02-16 11:38:49
#> 3 18632100 15360. 15346. 2026-02-16 11:02:20 2026-02-16 10:58:09
#> 4 21956100 13803. 248. 2026-02-16 15:59:17 2026-02-16 14:45:27
#> 5 5516100 15259. 0.0419 2026-02-16 15:12:19 2026-02-16 13:52:44
#> 6 6533100 12118. 0.0419 2026-02-16 15:59:17 2026-02-16 14:52:21
#> 7 7879100 15363. 0.0419 2026-02-16 14:41:08 2026-02-16 13:26:33
#> # ℹ 11 more variables: max_dist_gap <dbl>, max_t_gap <dbl>,
#> # max_dist_gap_id <chr>, max_t_gap_id <chr>, trip_distance <dbl>,
#> # duration <dbl>, dist_ok <lgl>, dur_ok <lgl>, dist_gap_ok <lgl>,
#> # t_gap_ok <lgl>, all_ok <lgl>Most of these were removed because of a large gap in data. Let’s take a look at one, trip 21956100:
# Filter dataframe to our tirp & distances
plot_df <- c53_cleaned_subtrips %>%
filter(trip_id_performed == "21956100") %>%
filter((distance >= 11000) & (distance <= 14000))
# Create a plot
gaps_plot <- ggplot() +
# Plot the points
geom_line(data = plot_df,
aes(x = event_timestamp, y = distance),
linewidth = 2, color = "lightcoral") +
geom_point(data = plot_df,
aes(x = event_timestamp, y = distance),
size = 2, color = "firebrick4") +
# Format the plot
theme_minimal() +
labs(x = "Time",
y = "Distance (m)",
title = "Gap on C53",
subtitle = "Trip 21956100")
gaps_plot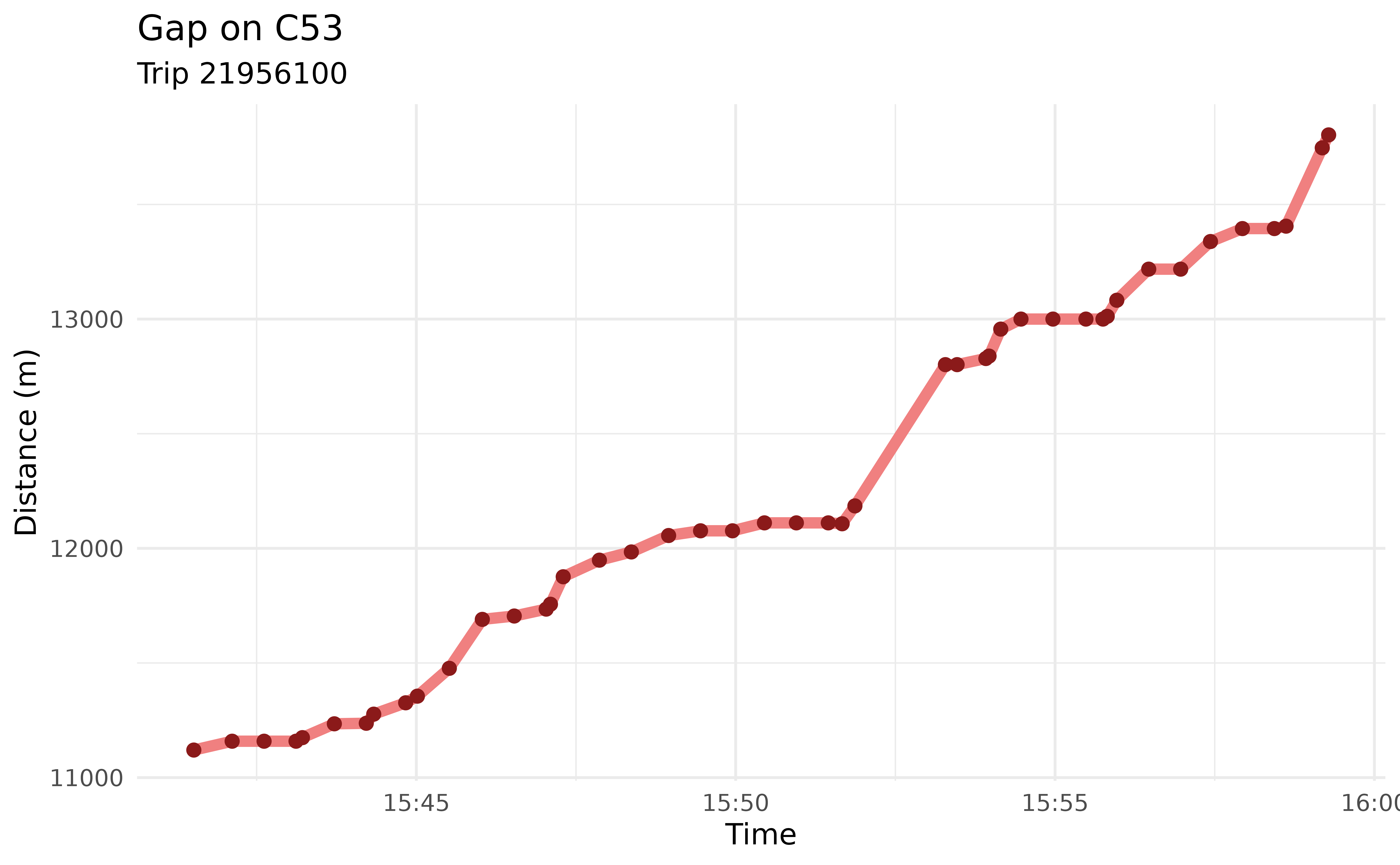
We can see this trip’s gap around 12,500 meters. The slope of this line is a bit steep, but is not unreasonable; it looks like a few GTFS-rt pings just went missing here. When we go to reconstruct a trajectory curve later, it may not be reasonable to interpolate between these points over such a large distance, so we’ll leave this trip out of our future analyses.
Step 6: Trim Trip Tails
Earlier in this vignette, we used a spatial buffer to clean what
appeared to be deadheads. But what if a trip deadheads close to – or
along – the route alignment? In Step 6, the function
trim_trips() is designed to handle these scenarios by
trimming the tails off of trips.
The function identifies the observations with the minimum and maximum distance values, and removes any observations which occur before/after these points. There is one main decision variable here: should beginning tails be trimmed, ending tails, or both? For this example, we’ll trim both ends of trips:
Running the Code
Let’s run trim_trips(), trimming both the beginning and
ends of each trip:
# Set parameters
c53_trim_type <- "both"
# Run function
c53_trimmed <- trim_trips(distance_df = c53_cleaned_incompletes,
trim_type = c53_trim_type)Exploring the Results
Let’s see what was removed:
# Pull dimensions
step6_obs <- dim(c53_trimmed)[1]
# Print
cat("Initial: ", step5_obs, " obs",
"\nAfter: ", step6_obs, " obs",
"\nDifference: ", (step5_obs - step6_obs), " obs removed")
#> Initial: 4089 obs
#> After: 4086 obs
#> Difference: 3 obs removedFor this example, it looks like there were no long deadheads along the route alignment. Let’s take a look at a the points removed:
c53_step6_removals <- trim_trips(
# Same settings as before
distance_df = c53_cleaned_incompletes,
trim_type = c53_trim_type,
# Return removals
return_removals = TRUE
)
print(c53_step6_removals)
#> # A tibble: 3 × 10
#> trip_id_performed event_timestamp distance min_dist_index max_dist_index
#> <chr> <dttm> <dbl> <int> <int>
#> 1 13478100 2026-02-16 15:59:11 2447. 1 88
#> 2 35294100 2026-02-16 11:26:04 18.8 3 202
#> 3 35294100 2026-02-16 11:26:34 94.3 3 202
#> # ℹ 5 more variables: row_index <int>, before_min <lgl>, after_max <lgl>,
#> # obs_ok <lgl>, location_ping_id <chr>We’ll plot the points removed along trip 35294100:
# Filter dataframe to our tirp & distances
plot_df <- c53_cleaned_incompletes %>%
filter(trip_id_performed == "35294100") %>%
filter(distance <= 200) %>%
# Join removals
left_join(y = (c53_step6_removals %>% select(location_ping_id, obs_ok)),
by = "location_ping_id") %>%
mutate(obs_ok = tidyr::replace_na(obs_ok, TRUE))
# Create a plot
trimmed_plot <- ggplot() +
# Plot the points
geom_point(data = plot_df,
aes(x = event_timestamp, y = distance,
color = obs_ok, shape = obs_ok),
size = 3, stroke = 3) +
# Format the points
scale_color_manual(name = "Point OK?",
values = c("FALSE" = "firebrick",
"TRUE" = "darkgreen")) +
scale_shape_manual(name = "Point OK?",
values = c("FALSE" = 4,
"TRUE" = 16)) +
# Format the plot
theme_minimal() +
labs(x = "Time",
y = "Distance (m)",
title = "Trimmed Trips on C53",
subtitle = "Trip 35294100")
trimmed_plot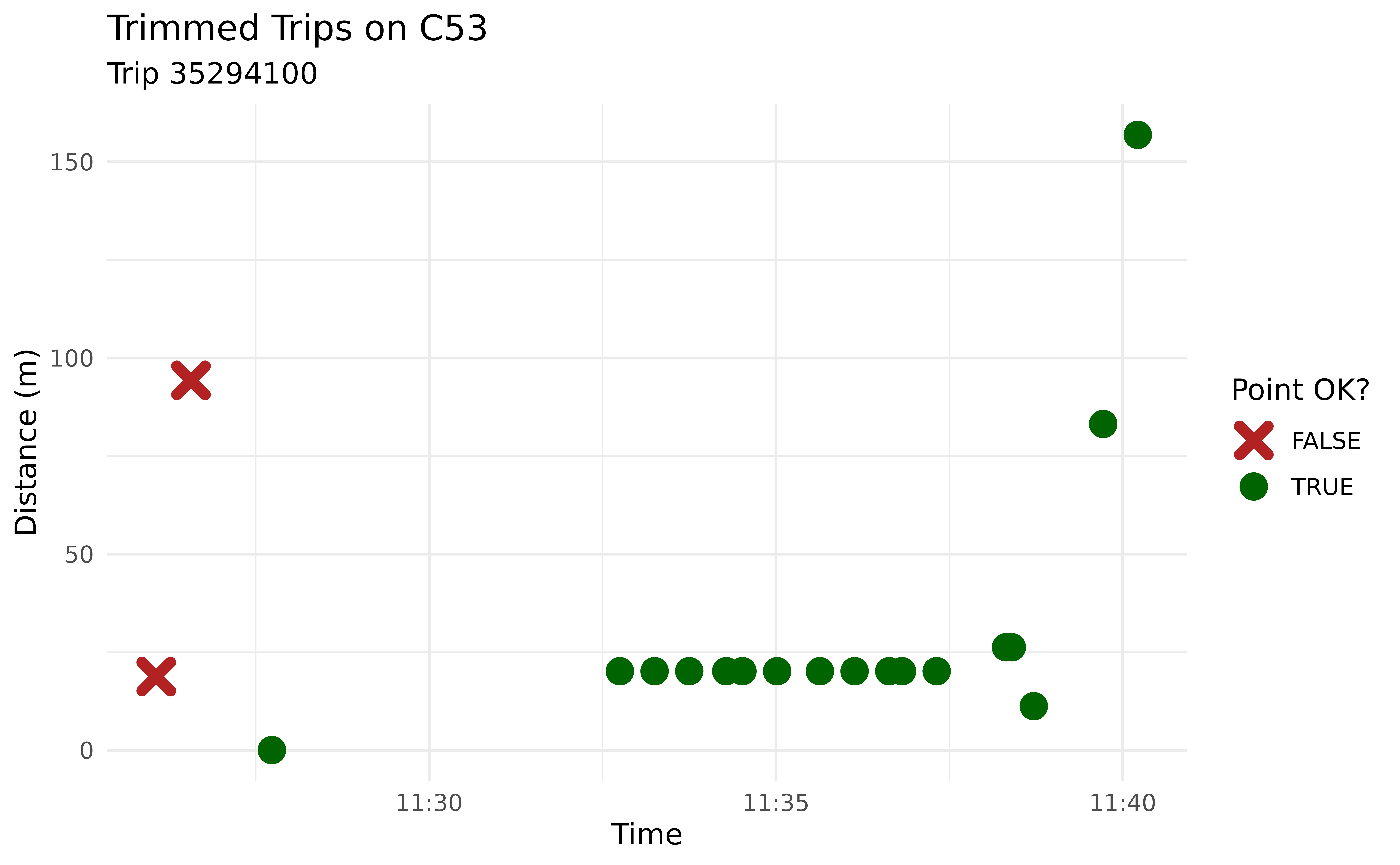
As noted before, this does not seem to be a long deadhead; just some
noise at the beginning of the route (by default,
clean_jumps() does not look for jumps at the beginning/end
of trips before a complete median window can be formed). Whether this
step is truly necessary will depend on your dataset.
Step 7: Correct for Monotonicity
While GPS noise can result in the large jumps we saw previously, it much more often causes small deviations in observed locations. This creates problems when fitting an interpolating trajectory curve, because if any points drift backwards, the curve will be neither monotonic nor invertible. In Step 7 (the last one!), we will “pull” any backtracking points up to where they should be. Optionally, data can also be made strictly monotonic.
The function make_monotonic() has two decision
variables:
Should speeds be corrected to satisfy monotonicity? AVL speed location can help fit an excellent interpolating curve, but the values of observed speeds must meet certain conditions (known as Fritsch-Carlson constraints) to produce a monotonic spline. If your AVL data has speed information, and you plan to use it when fitting a spline (the recommended interpolation method), set
correct_speed = TRUEto guarantee the fit is monotonic.Should the trajectory be made strictly monotonic? This will identify perfectly flat regions can give them a slight upward slope. To be invertible, a trajectory must be strictly increasing, and never constant.
Our AVL dataset does have speeds, and we do want an invertible and
monotonic final trajectory. As such, we’ll correct the speeds to meet
the Fritsch-Carlson constraints, and we will add a distance error of
0.001 meters (1 mm). More information is available at
help(make_monotonic).
Running the Code
Let’s run make_monotonic() using the parameters
discussed above:
# Set parameters
c53_dist_error <- 0.001
c53_correct_speeds <- TRUE
# Run function
c53_mono <- make_monotonic(distance_df = c53_trimmed,
correct_speed = c53_correct_speeds,
add_distance_error = c53_dist_error)Exploring the Results
This function modifies existing data, but does not remove any points. The total number of observations should stay the same:
# Pull dimensions
step7_obs <- dim(c53_mono)[1]
# Print
cat("Initial: ", step6_obs, " obs",
"\nAfter: ", step7_obs, " obs",
"\nDifference: ", (step6_obs - step7_obs), " obs removed")
#> Initial: 4086 obs
#> After: 4086 obs
#> Difference: 0 obs removedWe can check, though, if our dataset is now monotonic using
validate_monotonicity(). We’ll ask the function to validate
speeds as well:
# Trimmed DF
step6_val <- validate_monotonicity(distance_df = c53_trimmed,
check_speed = TRUE)
print(step6_val)
#> weak strict speed
#> FALSE FALSE FALSE
# Monotonic-corrected DF
step7_val <- validate_monotonicity(distance_df = c53_mono,
check_speed = TRUE)
print(step7_val)
#> weak strict speed
#> TRUE TRUE TRUEWe can see the trimmed dataset did not satisfy weak, strict, or
Fristch-Carlson speed conditions for monotonicity, but the corrected
dataset did. We can also see exactly which points were adjusted, and by
how much, using the parameter return_changes. Below we plot
an example trip, 21499100, around one stop it makes:
# Set filter parameters
plot_trip <- "21499100"
plot_dists <- c(13700, 13950)
# Get old DF
plot_df_before <- c53_trimmed %>%
filter(trip_id_performed == plot_trip) %>%
filter((distance >= plot_dists[1]) & (distance <= plot_dists[2])) %>%
mutate(speed_label = paste(round(speed, 1), "m/s", sep = ""))
# Get corrected DF
plot_df_after <- c53_mono %>%
filter(trip_id_performed == plot_trip) %>%
filter((distance >= plot_dists[1]) & (distance <= plot_dists[2])) %>%
mutate(speed_label = paste(round(speed, 1), " m/s", sep = ""))
# Plot
mono_plot <- ggplot() +
geom_point(data = plot_df_before,
aes(x = event_timestamp, y = distance,
color = "Uncorrected"),
size = 4, alpha = 0.6) +
geom_point(data = plot_df_after,
aes(x = event_timestamp, y = distance,
color = "Corrected"),
size = 3, alpha = 1) +
geom_label(data = plot_df_before,
aes(x = event_timestamp, y = distance,
color = "Uncorrected", label = speed_label),
nudge_y = -15, size = 2.5, show.legend = FALSE) +
geom_label(data = plot_df_after,
aes(x = event_timestamp, y = distance,
color = "Corrected", label = speed_label),
nudge_y = 15, size = 2.5, show.legend = FALSE) +
scale_color_manual(name = "Correction",
values= c("Uncorrected" = "indianred",
"Corrected" = "darkgreen")) +
# Format the plot
theme_minimal() +
labs(x = "Time",
y = "Distance (m)",
title = "Monotonic Correction on C53",
subtitle = paste("Trip ", plot_trip, sep = ""))
mono_plot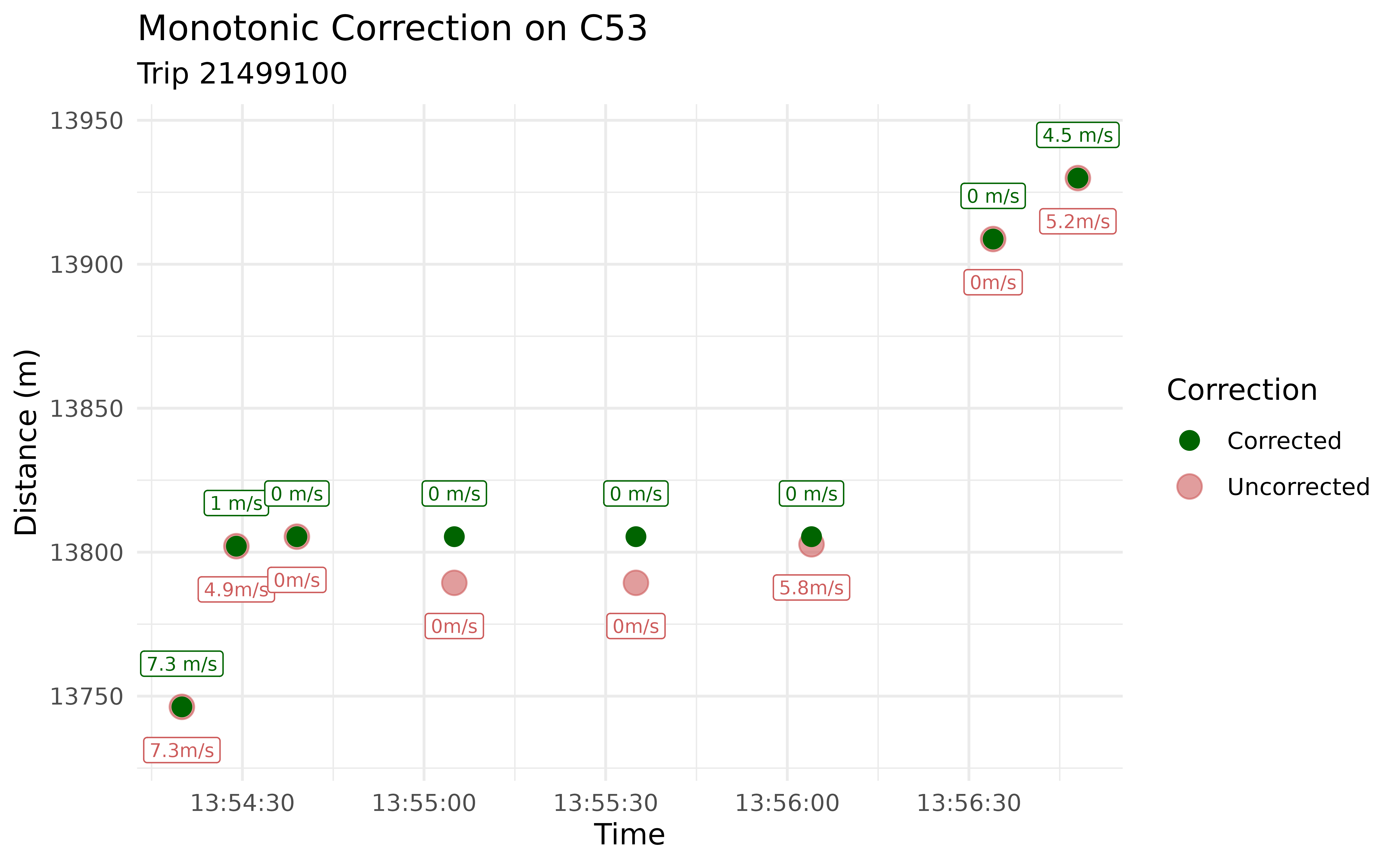
In this trip, the GPS backtracks slightly while the bus is supposed
to be stopped. The speeds are also larger than is physically feasible
for a monotonic trajectory as the bus enters and leaves this stop.
make_monotonic() identified and corrected both issues.
Conclusion
That was a lot of cleaning! But we now have a dataset free of
outliers and deadheading trips, and that we know will produce a
monotonic and invertible trajectory function. In the next vignette
(vignette("articles/intro-trajectories")) we will fit and
explore these interpolating curves.