Introduction to Trajectories
Source:vignettes/articles/intro-trajectories.Rmd
intro-trajectories.RmdIntroduction
In the previous vignette
(vignette("articles/data-workflow")), we saw how we can use
transittraj to clean our AVL data. We took care of
outliers, deadheading trips, noise, and non-monotonic observations. In
this vignette, we’ll apply the cleaned data (c53_mono) to
fit a trajectory function.
Let’s begin by loading the libraries we’ll be using:
Fitting a Trajectory Curve
Our ultimate goal is to fit an interpolating curve describing the
position of a transit vehicle at any point in time. Ideally, we could
fit an inverse curve, giving us the time the transit vehicle passes any
point in space. We can do both using
get_trajectory_fun().
transittraj supports a handful of methods for fitting
these functions. The simplest is linear interpolation without an
inverse. For more fine-grained analyses, though, we recommend fitting a
velocity-informed piecewise cubic interpolating polynomial.
This uses the speeds and distances, corrected for monotonicity, to fit a
cubic spline between each observation. This is the type of curve that
get_trajectory_fun() will fit by default
(interp_method = "monoH.FC" and
use_speeds = TRUE).
Using the data we cleaned in the previous vignette, let’s fit our trajectory functions:
# Run function
c53_traj <- get_trajectory_fun(distance_df = c53_mono,
interp_method = "monoH.FC",
use_speeds = TRUE,
find_inverse_fun = TRUE)transittraj stores the fit curves in a special object
class. This object stores a list of fit trajectories, one for each trip,
as well as the time and distances ranges for each trip. We can use
summary() to take a look inside the object:
summary(c53_traj)
#> ------
#> AVL Group Trajectory Object
#> ------
#> Number of trips: 24
#> Total distance range: 0 to 15370.75
#> Total time range: 1771257490 to 1771275553
#> ------
#> Trajectory function present: TRUE
#> --> Trajectory interpolation method: monoH.FC
#> --> Maximum derivative: 3
#> --> Fit with speeds: TRUE
#> Inverse function present: TRUE
#> --> Inverse function tolerance: 0.01
#> ------Interpolating
How do you use the fit curve to actually interpolate at new points?
We recommend using predict(), as this will ensure that the
curves aren’t used to extrapolate beyond the range of each trip. Using
predict(), there are three main ways we can interpolate:
retrieve distance values from times, retrieve time values from
distances, or retrieve time & distance pairs over a spatial
range.
Interpolating for Distance from Time
Let’s say you want to know where every vehicle is at a certain point
in time. We can do that by providing new_times to
predict(). Let’s see below:
# Run interpolating function
c53_time_interp <- predict(
object = c53_traj,
new_times = c(1771265000, 1771275000)
)
# Print full results
print(c53_time_interp)
#> event_timestamp trip_id_performed interp
#> 1 1771265000 1306100 8933.8531
#> 2 1771265000 18298100 10899.0966
#> 3 1771265000 21499100 4006.3031
#> 4 1771265000 21555100 13912.1101
#> 5 1771265000 22663100 6944.8919
#> 6 1771275000 10185100 2883.3087
#> 7 1771275000 10249100 7986.3226
#> 8 1771275000 13478100 140.0436
#> 9 1771275000 3597100 6792.4405Here, interp will be the distance in meters from the
route’s beginning. You’ll notice that, even though we have 24 trips,
there were only four to five distance for each timepoint. This is
because predict() will only interpolate a distance value
for trips that were actually running at that point in time.
Using a similar function call, we can also find the speed of the
vehicle at any point in time by setting the deriv parameter
in predict():
# Run interpolating function
c53_speed_interp <- predict(
object = c53_traj,
new_times = c(1771265000, 1771275000),
deriv = 1
)
# Print results
print(c53_speed_interp)
#> event_timestamp trip_id_performed interp
#> 1 1771265000 1306100 1.675073e-01
#> 2 1771265000 18298100 1.464849e+00
#> 3 1771265000 21499100 1.372977e+01
#> 4 1771265000 21555100 6.424041e-01
#> 5 1771265000 22663100 1.508816e+00
#> 6 1771275000 10185100 4.473051e-01
#> 7 1771275000 10249100 4.354498e-05
#> 8 1771275000 13478100 7.213864e-01
#> 9 1771275000 3597100 9.079446e-01Here, interp will be the speed in meters per second.
Finding speeds requires starting from time values; we cannot get speeds
from distance values.
Interpolating for Time from Distance
One of the most common applications of the fit trajectory curve is to
find the time at which each vehicle passed a point along its route. To
do this, we’ll use predict() with the
new_distances parameter. We’ll begin by finding the
distance of each stop along the route using
get_stop_distances():
# First, use stop_times to find which stop_ids are timepoints
c53_timepoints <- c53_gtfs$stop_times %>%
distinct(stop_id, timepoint)
# Now, find stop distances and join the timepoints column
c53_stops <- get_stop_distances(gtfs = c53_gtfs,
shape_geometry = c53_shape,
project_crs = dc_CRS) %>%
# Join timepoint info to each stop ID
left_join(y = c53_timepoints,
by = "stop_id") %>%
# Polish up the result
select(-c(shape_id, stop_code, stop_desc,
zone_id, stop_url)) %>%
mutate(timepoint = if_else(condition = (timepoint == 1),
true = "Yes",
false = "No"))
# Print header
head(c53_stops)
#> # A tibble: 6 × 4
#> stop_id stop_name distance timepoint
#> <chr> <chr> <dbl> <chr>
#> 1 2584 Alabama Av SE+15 Pl SE 677. No
#> 2 2609 Alabama Av SE+Stanton Rd SE 880. No
#> 3 2683 Alabama Av SE+18 Pl SE 1155. No
#> 4 2793 Alabama Av SE+22 St SE 1605. No
#> 5 2811 Alabama Av SE+24 St SE 1807. No
#> 6 2867 Alabama Av SE+Jasper St SE 2037. NoNow that we have some distances, let’s interpolate using
predict():
# Run interpolating function
c53_stop_crossings <- predict(
object = c53_traj,
new_distances = c53_stops
)
# Print header
head(c53_stop_crossings)
#> # A tibble: 6 × 6
#> stop_id stop_name distance timepoint trip_id_performed interp
#> <chr> <chr> <dbl> <chr> <chr> <dbl>
#> 1 2584 Alabama Av SE+15 Pl SE 677. No 10185100 1.77e9
#> 2 2584 Alabama Av SE+15 Pl SE 677. No 10249100 1.77e9
#> 3 2584 Alabama Av SE+15 Pl SE 677. No 1306100 1.77e9
#> 4 2584 Alabama Av SE+15 Pl SE 677. No 13437100 1.77e9
#> 5 2584 Alabama Av SE+15 Pl SE 677. No 13478100 1.77e9
#> 6 2584 Alabama Av SE+15 Pl SE 677. No 1699100 1.77e9Now we have the crossing time, labeled interp at each
stop for each trip. The interpolated times are in seconds of epoch
time.
Interpolating for Time & Distance Pairs Over a Range
The final interpolation method allows you to specify a range of
distances, and a timestep over which to interpolate within this range.
Here, transittraj will use your trajectory’s inverse
function to find the time each trip enters and exits the
distance_lims, then interpolate every timestep
seconds that the vehicle stays in that range.
To see what this does, let’s interpolate some timepoints for all trips over U Street between 13th and 14th Streets NW. We’ll begin by pulling a vector of this distance range:
# Get distance limits of U St between 13th and 14th
U_St_lims <- c53_stops %>%
filter(stop_name %in% c("U St NW+13 St NW",
"U St NW+14 St NW")) %>%
pull(distance)
print(U_St_lims)
#> [1] 12800.72 13000.01Next, we can put this into predict() using the
distance_lims parameter, alongside a timestep
of 1 second:
# Run interpolating function
c53_USt_interp <- predict(
object = c53_traj,
distance_lims = U_St_lims,
timestep = 1
)
# Print header
head(c53_USt_interp)
#> # A tibble: 6 × 3
#> trip_id_performed event_timestamp interp
#> <chr> <dbl> <dbl>
#> 1 1115100 1771258405. 12801.
#> 2 1115100 1771258406. 12803.
#> 3 1115100 1771258407. 12805.
#> 4 1115100 1771258408. 12808.
#> 5 1115100 1771258409. 12811.
#> 6 1115100 1771258410. 12815.We can see that, for the printed trip, the first timepoint occurs at
the beginning of U_St_lims, then
event_timestamp increments one second per row afterwards.
The interp column is the distance at each time (you can
also set deriv here). To better understand see what this
did, we’ll generate a plot of these generated points. Below, we first
“center” each trip to start at 0 seconds, then plot point colored by
trip:
# "Center" all trips to start at 0 time
c53_USt_centered <- c53_USt_interp %>%
group_by(trip_id_performed) %>%
mutate(event_timestamp = event_timestamp - min(event_timestamp)) %>%
rename(distance = interp)
# Create plot
USt_plot <- ggplot(data = c53_USt_centered) +
# Add points
geom_point(aes(x = event_timestamp, y = distance,
color = trip_id_performed),
size = 2, alpha = 0.4) +
# Color points by trip
scale_color_viridis_d(guide = "none") +
# Theming
theme_minimal() +
labs(x = "Time (s)",
y = "Distance (m)",
title = "C53 Second-by-Second Position",
subtitle = "U St NW between 13th and 14th St NW")
USt_plot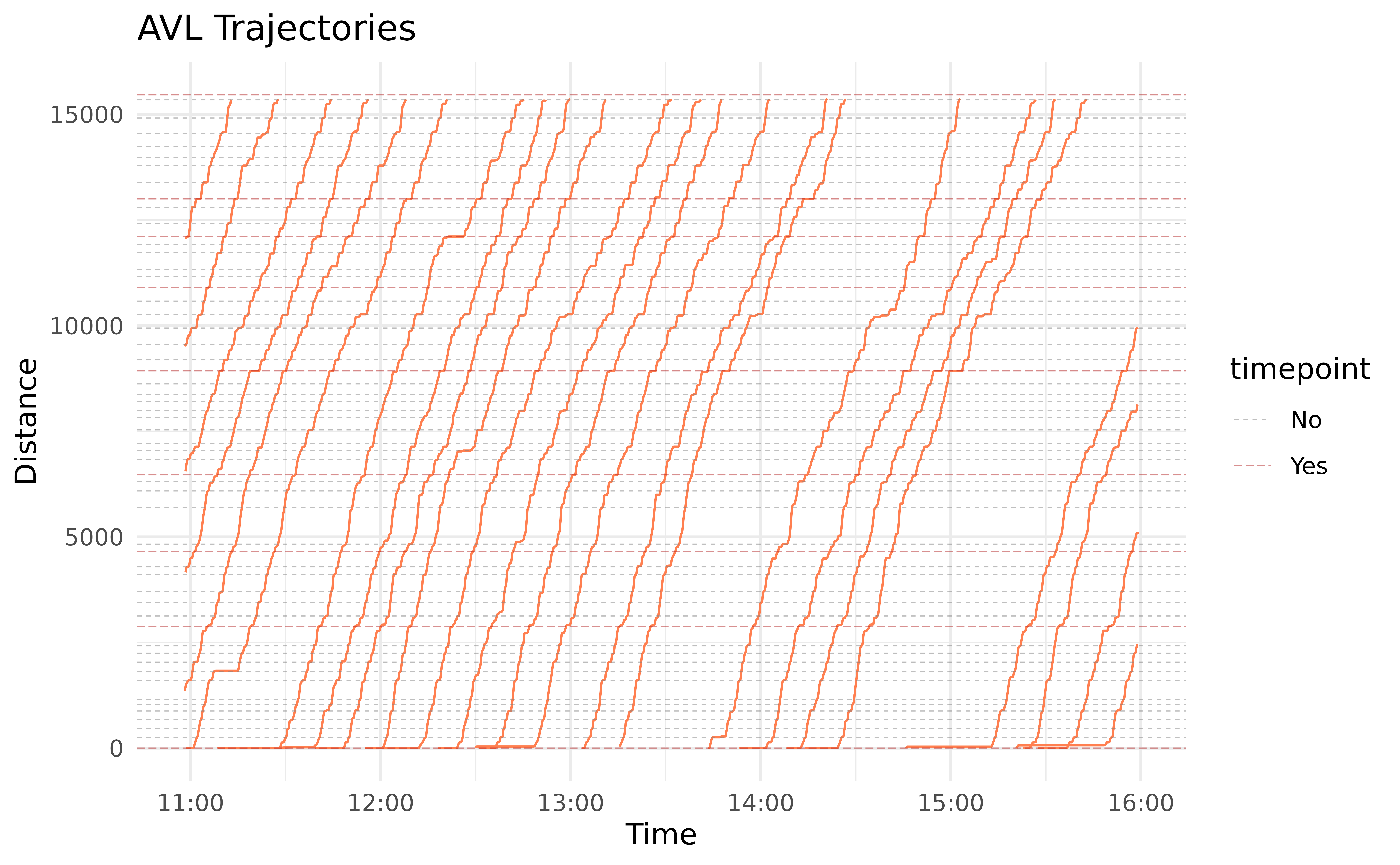
You could retrieve identical results by giving predict()
a new_times sequence spanning the range of the trajectory’s
event_timestamp’s, then filtering to the desired distance
range. For large datasets – spanning, for example, months –, however,
this would require a massive sequence. If an inverse function
is available, using distance_lims and timestep
is a much more efficient way to generate high-resolution trajectory
profiles for a large number of trips, especially if you are interested
in studying a specific region in space.
Visualizing Trajectories
Quick Plots
Now its time for the fun part – plotting our trajectory curves. We
can use plot() to easily generate a plot of all
trajectories:
plot(c53_traj)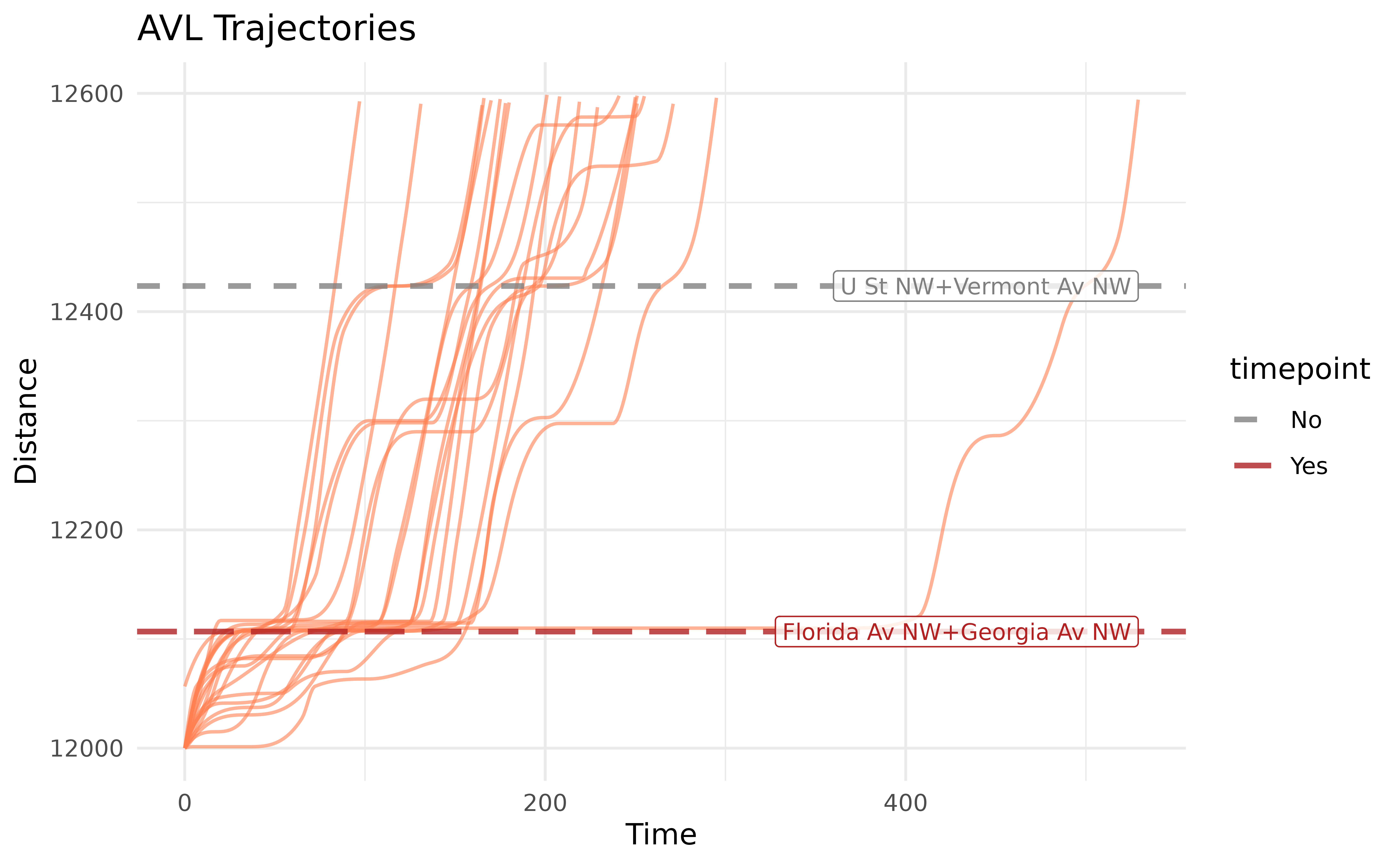
plot() is intended for quick visualizations of
trajectories, and as such does not allow for much customization. In the
next section, we’ll use plot_trajectory() to create more
interesting plots.
Detailed Trajectories
For more customization, we recommend using
plot_trajectory(). In addition to a trajectory object, you
can add a dataframe of feature distances, such as the
c53_stops dataframe we made earlier. Most layer aesthetics
can be controlled using input parameters. For features and trajectories,
the linetypes and colors can also be mapped to attributes of that
specific layer using a dataframe:
# Set formatting options for C53 stops
stop_formatting <- data.frame(timepoint = c("Yes", "No"),
color = c("firebrick", "grey50"),
linetype = c("longdash", "dashed"))For mapping dataframes, at least one column must match a column in
the layer being mapped to. The other columns must be color
and/or linetype, telling transittraj which
feature they describe.
We can plug all that in to plot_trajectory() to generate
our formatted plot:
# Run plotting function
traj_plot <- plot_trajectory(
# Provide input data
trajectory = c53_traj,
feature_distances = c53_stops,
# Format features
feature_color = stop_formatting,
feature_type = stop_formatting,
feature_width = 0.2, feature_alpha = 0.5,
# Format trajectories
traj_width = 0.4, traj_alpha = 1
)
traj_plot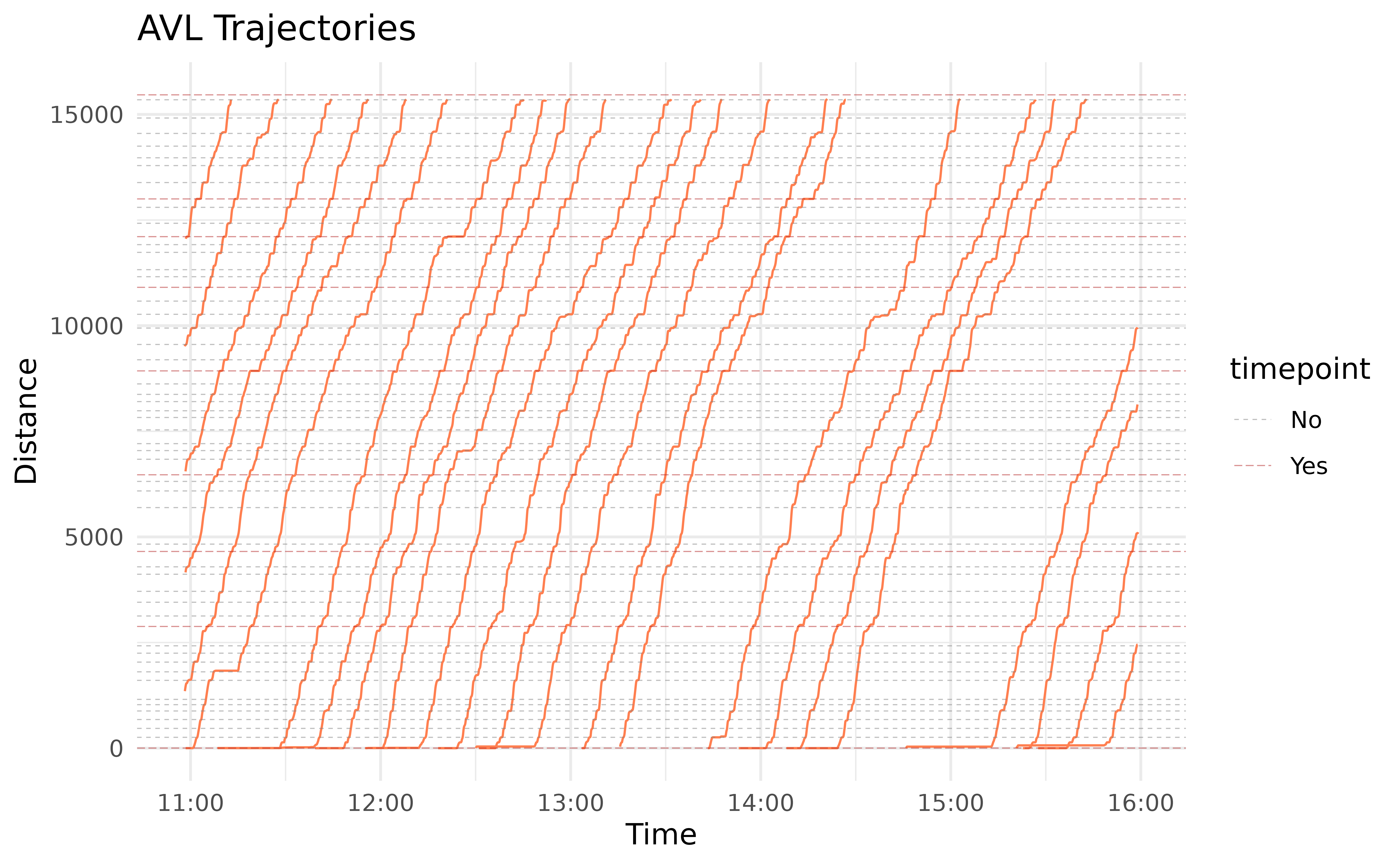
It’s hard to see what’s actually going on here. The benefits of the
cleaning we did, and of fitting a spline trajectory, become much more
apparent when we zoom in. Below we use the distance_lim
parameter to zoom into the intersection of Florida Ave & U St. This
is a large intersection with complex geometry and stops on either
side.
We’ll use two additional plotting parameters here. First,
center_trajectories will center each trajectory to start at
the same point in time. Second, label_field will create a
label on our feature lines using the specified field from
c53_stops.
# Set parameters
fl_U_intersection_lims <- c(12000, 12600)
# Run function
fl_U_plot <- plot_trajectory(
# Provide input data
trajectory = c53_traj,
feature_distances = c53_stops,
center_trajectories = TRUE,
distance_lim = fl_U_intersection_lims,
timestep = 1,
# Format fetures
feature_color = stop_formatting,
feature_type = stop_formatting,
feature_width = 1, feature_alpha = 0.8,
# Format trajectories
traj_width = 0.6, traj_alpha = 0.6,
# Add labels
label_field = "stop_name", label_pos = "right",
label_alpha = 0.8
)
fl_U_plot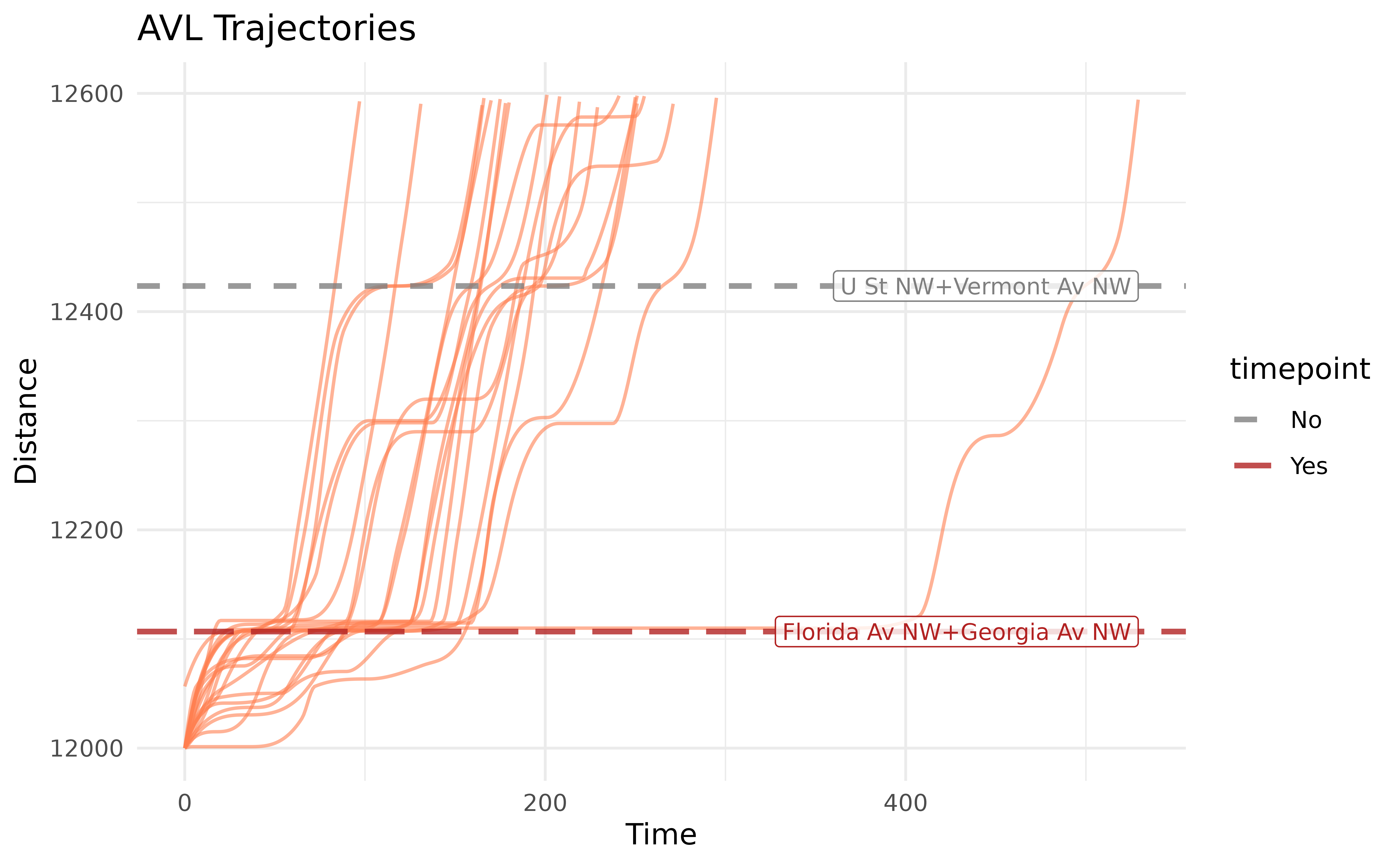
We can glean some insights from this. Almost every trip stops at Florida & Georgia, either to serve the stop or wait for the signal. One trip sits there for a particularly long time. A handful of others stop at the signal in between these two stops, and a couple more stop at U & Vermont. A few trips have slowdowns between these stops and signals, potentially due to congestion.
Check out help(plot_trajectory) for a full discussion of
the formatting features available.
Line Animations
Another fun way to visualize transit vehicle trajectories is to
animate them. Use plot_animated_line() to animate vehicles,
as points, moving along a straight line.
The formatting process works very similarly with
plot_animated_line() as it does with
plot_trajectory(). A dataframe can be used to map the
outline color and shape attributes of stop and
vehicle points to their attributes.
# Set parameters
stop_formatting <- data.frame(timepoint = c("Yes", "No"),
outline = c("red1", "grey30"),
shape = c(22, 21))For this plot, we’ll zoom in to the Florida Ave-U St corridor of the route. Now we can generate our line animation:
# Set distance limits
fl_U_corridor_lims <- c(9500, 15500)
# Run function
line_anim <- plot_animated_line(
# Add input data
trajectory = c53_traj,
feature_distances = c53_stops,
distance_lim = fl_U_corridor_lims,
timestep = 1,
# Format features
feature_outline = stop_formatting,
feature_shape = stop_formatting,
feature_size = 3, feature_stroke = 1.5,
# Add labels
label_field = "stop_name",
label_pos = "right", label_size = 3,
# Format route & vehicles
route_color = "indianred2",
veh_alpha = 0.9, veh_size = 4
)
line_animThe animation shows us that most trips stop primarily at their stops, either due to signals or to serve the stop. There are, though, occasional slow downs between these stops. You can even see that trip that sits at Florida & Georgia for a long time (at around 0:16 seconds).
You’ll also notice that we’ve uploaded this animation to YouTube and
embedded it in the vignette. We did this so we could produce a smooth,
high-resolution video that doesn’t need to be re-rendered every time
this vignette is built. By default, transittraj’s animation
functions will return a gif. Check out
gganimate::animate() for options to render videos.
Map Animations
The final visualization we’ll make is an animated map. The concept is similar to the animated line we saw above, but instead of simplifying the route, we’ll draw it spatially and show the vehicles traveling through the city.
The function plot_animated_map() has formatting and
feature options very similar to the previous two visualization
functions. We can reuse the formatting options from
plot_animated_line() here.
# Run function
map_anim <- plot_animated_map(
# Add trajectory, shape, & feature data
trajectory = c53_traj,
shape_geometry = c53_shape,
feature_distances = c53_stops,
# Format features
feature_outline = stop_formatting,
feature_shape = stop_formatting,
feature_size = 3, feature_stroke = 2,
# Format route
route_color = "indianred3", route_width = 4,
bbox_expand = 700,
# Format vehicles
veh_size = 6, veh_stroke = 3, veh_alpha = 0.9
)
map_animThis animation helps use see spatially where buses start to bunch
together, such as the two vehicles entering Florida at around 0:34
seconds, or the three vehicles near Florida at around 0:45 seconds.
distance_lims can be used to zoom in on specific regions,
just as before.
Conclusion
In this vignette we saw how we can easily fit an interpolating
trajectory curve to our cleaned AVL data. We used this to interpolate
for new time, distance, and speed points along the route. We also
explored some ways we can plot and visualize the trajectories. Future
vignettes (vignette("articles/indygo-signals")) will
explore real-world applications of trajectories.